Retooling the Translation Machine Could Expand the Chemical Repertoire of Cells
Synthetic biologists have become increasingly creative in engineering yeast or bacteria to churn out useful chemicals — from fuels to fabrics and drugs — beyond the normal repertoire of microbes.
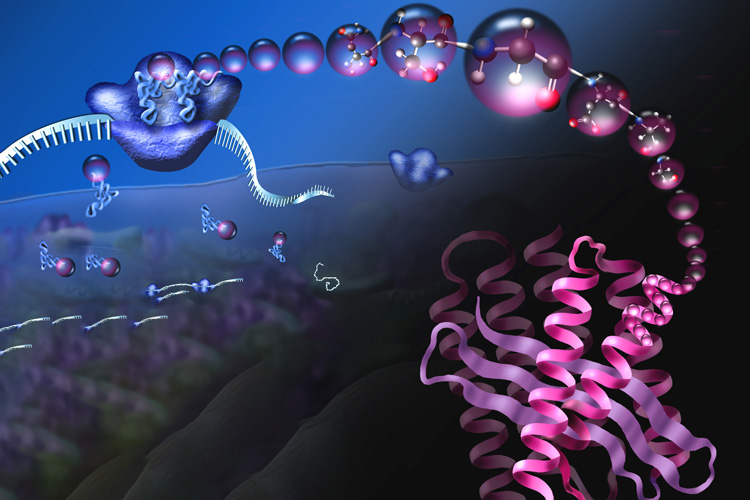
But a multi-university group of chemists has a more ambitious goal: to retool the cell’s polypeptide manufacturing plants — the ribosomes that spin amino acids into protein — to generate polymer chains that are more elaborate than what can now be made in a cell or a test tube.
The $20 million research enterprise centered at the University of California, Berkeley, is now reporting significant progress toward that goal, as evidenced by three new papers that tackle three major hurdles: how to reprogram cells to supply the ribosome with building blocks other than the alpha-amino acids that make up all proteins today; how to predict which building blocks make the best substrates; and how to tweak the ribosome to incorporate these novel building blocks into polymers.
The ultimate goal of the National Science Foundation Center for Genetically Encoded Materials (C-GEM) is to make the translation system fully programmable, so that introducing mRNA instructions into the cell along with new building blocks — not the alpha-amino acids found today — will allow the ribosome to produce an unlimited variety of new molecular chains. These chains could form the basis for new bio-materials, new enzymes, even new drugs.
The papers, appearing in the journals Nature Chemistry and ACS Central Science, are the beginning of a playbook for reengineering the cellular synthetic machinery to produce never-before-seen polymers, including bio-polymers and circular polymers, which are called peptide macrocycles, with predetermined or completely unforeseen applications.
“C-GEM is working to biosynthesize molecules that have never before been made in a cell and that are designed to have unique properties. The tools could be applied broadly by polymer chemists, medicinal chemists and biomaterials scientists to generate bespoke materials with new functions,” said C-GEM director Alanna Schepartz, the T.Z. and Irmgard Chu Distinguished Chair in Chemistry and professor of molecular and cell biology at UC Berkeley. “The ultimate goal is to expand the function and versatility of proteins and polypeptides, as both materials and pharmaceuticals.”
One example, she said, would be to program the ribosome to synthesize a polymer that is a cross between spider silk — one of the toughest natural proteins — and nylon, a polymer now made in chemical reaction chambers. While spider silk can now be made in genetically engineered microbes, the technology C-GEM is developing could allow similar microbes to make an infinite variety of polymers blending the building blocks of silk and nylon, all of them new to chemists and with unique properties. The technology could also be used to make protein-like polymers more resistant to heat than natural proteins.
A powerful aspect of a programmable ribosome machine that can synthesize polymers is that it allows researchers to evolve the polymers to perfect their activity, just as proteins have evolved over hundreds of millions of years to improve the fitness of cells and organisms.
“We’ve had protein polymers evolving on the planet for billions of years, but we’ve been restricted in what those polymers are because the building blocks are the same 20 amino acids,” said Jamie Cate, UC Berkeley professor of chemistry and of molecular and cell biology. “If we can develop a system where you can actually apply evolution to these new polymers, then it’s like a platform that anybody who has a creative idea can use to evolve a polymer to something they want.”
Such a system builds on the directed evolution of protein enzymes for which Frances Arnold, a UC Berkeley alumna, received the 2018 Nobel Prize in Chemistry.
“It’s a step beyond what Frances Arnold did in developing directed evolution,” Cate said. “She developed directed evolution for proteins. What we’re trying to do is set up a way that you could do this for polymers never before evolved in nature.”
Engineering an entirely new ribosome
In all cells, proteins are assembled by a nanomachine, the ribosome, that accepts instructions from an RNA molecule called messenger RNA (mRNA) — mRNA is akin to a working copy of a gene’s DNA code — and reads those instructions to assemble a protein, amino acid by amino acid. Amazingly, the linear protein chain almost always folds into a well-defined 3D structure, ready to serve its evolved purpose: as an enzyme to catalyze reactions in the cell, as a structural component of the cell, or as a regulator of other cellular activities.
Ten years ago, retooling this complex nanomachine seemed impossible. But Schepartz’s tenacity in pushing for a project to accomplish this goal resulted in C-GEM, which is three years into its first five-year funding cycle.
One of the center’s goals is to supply the ribosome with building blocks — so-called monomers — other than alpha-amino acids. To achieve this goal, the C-GEM team focused on the enzymes that load amino acid monomers onto transfer RNA (tRNA), the molecules that ferry amino acids to the ribosome. Each tRNA is bar-coded to indicate which of the 20 amino acids it carries.
As reported in a Nature Chemistry paper published June 1 and co-authored by Schepartz and graduate students Riley Fricke and Cameron Swenson, the team discovered that one family of tRNA synthetases could load tRNA with four different non-alpha-amino acids. One of these was a building block of various polyketide therapeutics, including the antibiotics erythromycin and tetracycline.
“We identified enzymes that load tRNAs with monomers that differ structurally from anything that has been loaded on a tRNA before,” Schepartz said. “One of the monomers is a precursor that could be used to assemble polyketide-like molecules. Scientists have been trying for decades to reengineer polyketide synthase modules to generate libraries of natural products. These studies have taught us a ton about the sophistication of these modules, but the engineering part has been very hard.”
The novel monomers were willingly accepted by the native ribosome in the bacteria E. coli, demonstrating that it’s possible to incorporate different types of chemistries into the normally all-amino acid protein polymer.
“Antibiotic resistance is an enormous problem,” she added. “If we could help solve that problem by generating novel molecules whose functions encode unique modes of action, that would be an enormous contribution.”
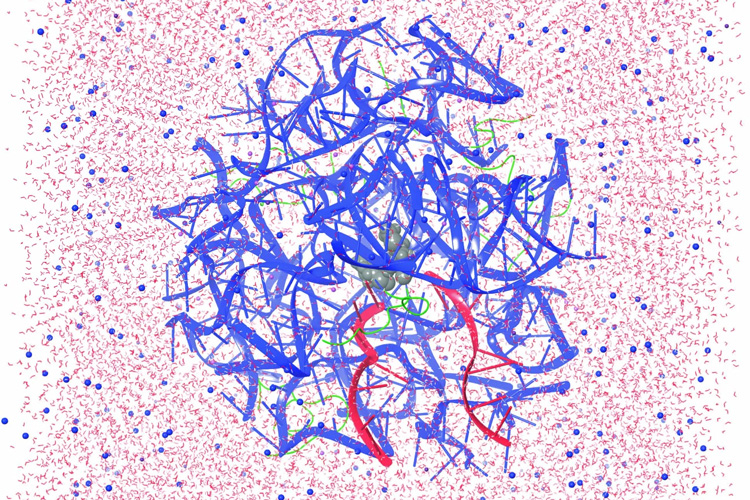
In a second paper, which appeared May 30 in ACS Central Science, lead author and postdoctoral fellow Chandrima Mujumdar, along with Cate and Schepartz, used cryogenic electron microscopy (cryo-EM) to obtain detailed structures of three related monomers — none of them alpha-amino acids — bound to the E. coli ribosome. The details show how these monomers bind — though much more poorly than do amino acids — and provide hints on how to alter the monomers or the ribosome to improve the ribosome’s ability to use them to build novel polymers.
In a third paper, which appeared June 12 in Nature Chemistry, Cate, Schepartz and lead author Zoe Watson, a postdoctoral fellow, report the cryo-EM structure of the E. coli ribosome while binding normal alpha-amino acids. For this paper, the team collaborated with the company Schrödinger Inc. of San Diego, which uses computers to model protein binding. Ara Abramyan of Schrodinger used the cryo-EM structure as a starting point to run metadynamic simulations to help understand which non-natural monomers will react in the ribosome’s catalytic center — the peptidyl transferase center (PTC) — and which will not.
Schepartz and Cate emphasized that all of these tweaks to the ribosomal system must work inside a living cell independently of the normal ribosomes so that the production of new polymers does not interfere with the day-to-day protein production necessary for life.
“We want enzymes — synthetases — and ribosomes that could be used in a cell, because that’s how this work will be scalable,” Schepartz said. “That goal requires robust ribosomes, great enzymes and a lot of understanding about the chemistry of how these complex molecular machines work. It’s a hard problem, but lots of fun. And we get to expose students and postdocs to some really great science.”
Among the authors are C-GEM investigators Matthew Francis of UC Berkeley, Scott Miller of Yale University, Abhishek Chatterjee of Boston College, Bhavana Shah and Zhonqi Zhang of Amgen Inc., and C-GEM managing director Sarah Smaga. Schepartz is also a member of the Chan Zuckerberg Biohub and the California Institute for Quantitative Biosciences (QB3). Cate is a member of the Innovative Genomics Institute. Both Schepartz and Cate are faculty scientists at Lawrence Berkeley National Laboratory.
NSF provided most of the funding under grant CHE 2002182.
RELATED INFORMATION
- Expanding the substrate scope of PylRS enzymes to include non-⍺-amino acids in vitro and in vivo (Nature Chemistry)
- Atomistic simulations of the E. coli ribosome provide selection criteria for translationally active substrates (Nature Chemistry)
- Aminobenzoic acid derivatives obstruct induced fit in the catalytic center of the ribosome (ACS Central Science)
