Nano-sized sensors learn new biological tricks
Christopher Jackson, a graduate student in the Landry Lab at QB3-Berkeley, explores how a better understanding of nanotechnology interactions with biological systems can improve neuroimaging and COVID-19 testing.
In spite of the tremendous advances in modern medicine, there are still mysteries about routine processes in the human body that continue to elude scientists. For example, researchers have long wondered how common neurochemicals, such as dopamine and serotonin, work inside the brain to create human emotions. These same chemicals play a critical role in diseases such as addiction, Huntington’s disease, and Parkinson’s disease, for which there are only limited treatments. Why? Until now, the tools didn’t exist to take measurements at the very small time and length scales at which these chemical processes occur.
Markita Landry, a Professor of Chemical and Biomolecular Engineering at the University of California, Berkeley, is fascinated by these biological processes that occur at the nano-scale. “A lot of the more recent developments in sensor technology have been in attaining greater levels of spatial resolution for imaging biochemistry and specifically neurochemistry,” says Landry. Nanotechnologies, and in particular nanosensors, are the tools that have enabled these advances. However, the effectiveness of these sensors dwindles when used in messy biological environments such as saliva or blood, a necessary hurdle that must be overcome for neuroimaging or other applications such as COVID-19 testing.
Carbon Nanotubes: a Scaffold for Nanosensor Probe Development
The Landry lab’s tools of choice are single-walled carbon nanotubes (SWNTs), a nanomaterial that is uniquely small, semiconducting, and tough. SWNTs are extremely rigid, high aspect ratio cylinders of carbon typically around 1 nanometer in diameter by 500 nanometers in length—about 10,000 times thinner than a human hair. This distinct shape makes them well equipped to pass through otherwise impenetrable barriers that other nanomaterials struggle with such as the cell membrane. SWNTs also have an extremely large surface area that the Landry lab can take advantage of to chemically attach molecules with different functions, from increasing the sensor stability to targeting certain locations in the cell.
Besides their unique mechanical properties, the SWNTs that the Landry lab uses have another key attribute that make them ideal sensor candidates – they’re semiconductors that emit light in the near-infrared (nIR) light spectrum. Unlike visible light, near-infrared light can penetrate through skin, tissue, blood, and water with minimal absorbance and scattering, allowing researchers to visualize otherwise invisible environments inside the body.
A sensor, in its simplest form, is composed of two parts: one that recognizes the analyte you want to detect and one that is a signal transducer, converting that recognition into something that researchers can measure. When some molecule binds to a SWNT surface, “there’s a change in the carbon nanotube fluorescence – either an increase or decrease in the signal or a shift in the wavelength of emission,” says Rebecca Pinals, a 5th-year graduate student in the Landry lab. The tunability of their SWNT system has allowed the Landry lab to quickly develop sensors for a wide variety of molecules, including neurochemicals that were impossible to visualize just a few years ago.
When she first arrived at UC Berkeley, Landry turned her focus to the brain, which she describes as “a uniquely challenging place to take measurements.” In particular, the lab’s early work focused on developing a particular sequence of DNA that is sensitive to catecholamines, a class of neurotransmitters including dopamine and norepinephrine. When attached to carbon nanotubes and injected into mouse brain slices, these nanosensors can detect fast fluctuations of dopamine levels in the area between neurons. While the selection of a DNA sequence was initially a more luck-based screening approach, recent work has focused on evolving DNA sequences for improved recognition through cycles of screening and selection for more rational sensor design.
Pinals walks us through the process of making one of these sensors in the lab. First, the DNA (or another recognition polymer) is attached to the SWNT, either via probe tip sonication or with different types of covalent chemistry. “We use this surface functionalization not only to make a sensor, but also to help solubilize the carbon nanotubes in water,” explains Pinals, and additional types of molecules, such as surfactants, may be added to improve solubility. These sensors are then mixed with the analyte of interest, such as dopamine, in a controlled well-plate and their fluorescence response is measured via spectroscopy. Once the most responsive candidates have been identified, those SWNT sensors can be injected into mouse brain slices for ex vivo testing.
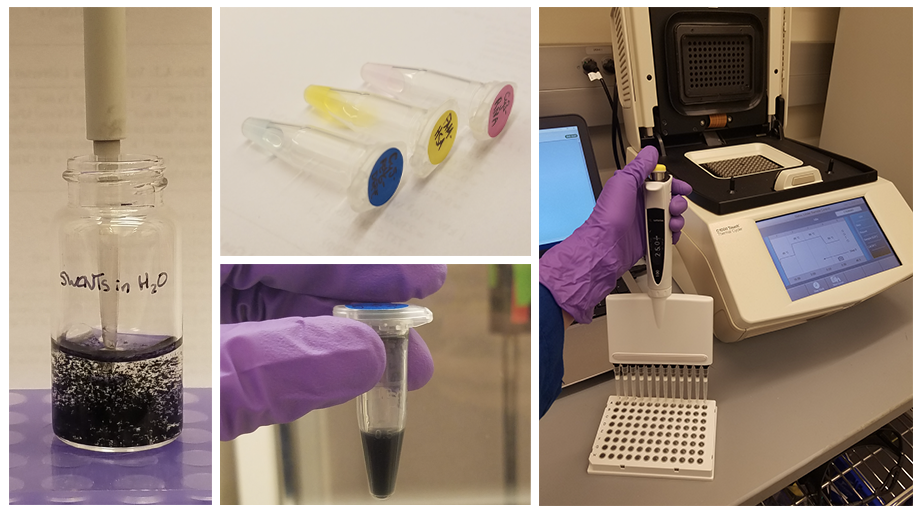

From Benchtop to Biology
Early on, the lab realized that just because these SWNT sensors work for pure samples in a clean benchtop environment, it doesn’t mean that those results will translate to biological environments like the brain or blood. “The translation from in vitro characterization to in vivo use is one of the largest challenges in the field,” emphasizes Landry, and figuring out why that happens likely isn’t a single answer. Pinals elaborates further: “when you inject these nanosensors into the body, they encounter proteins, lipids, the extracellular matrix, and other complex systems.” During this process, biomolecules from the surrounding media rapidly adsorb to exposed SWNT surfaces, forming a “protein corona.”
This phenomenon isn’t unique to SWNTs or even nanomaterials in general. For example, this process is occurring right now with the food in your refrigerator, but we don’t notice because it happens at such a low surface area to volume ratio. “When we miniaturize everything, the surface area to volume ratio becomes quite significant,” explains Landry. For the SWNT sensors, the high surface area that allows for robust tunable functionality can now become a problem.
Corona formation, and in particular biofouling, where nonselective, unpredictable proteins stick to the SWNT surface, can lead to sensor failure. Pinals’ graduate career has centered on developing new techniques to understand protein coronas, which depend on a wide range of nanoparticle properties, such as hydrophobicity, hydrophilicity, electrostatic charge, and the biological environment they’re in. Until recently, there were limited tools that could identify what kinds of proteins actually make up the corona or provide any insight into its formation mechanism. To overcome this challenge, Pinals and labmate Darwin Yang developed an assay capable of measuring the real-time binding of proteins onto the SWNT surface in solution.
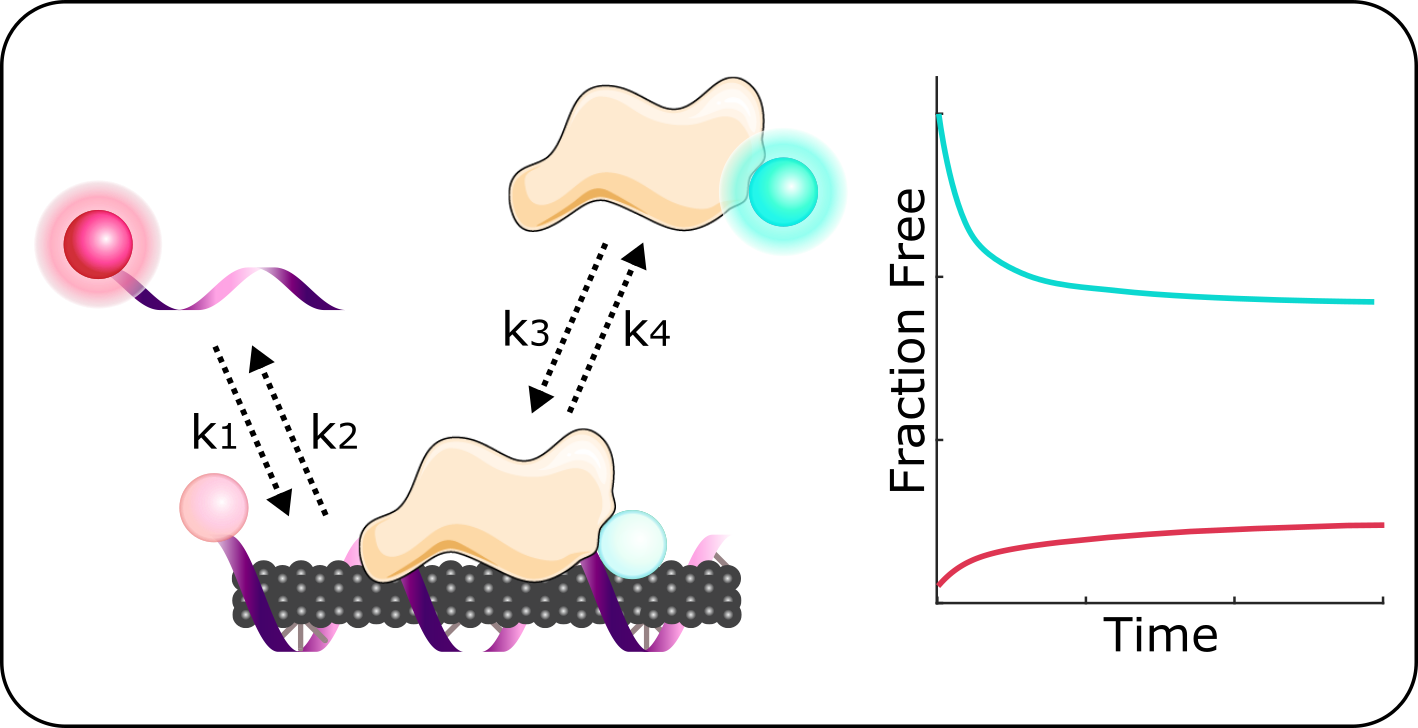
“We want to measure the amount of the DNA on the carbon nanotube surface, then track its exchange with proteins that are floating around in solution in real time,” says Pinals. To achieve this, it first requires labeling both the DNA and proteins with two distinct fluorophores, molecules that emit light at distinct wavelengths and can be measured via spectroscopy. When these fluorophore-labeled molecules are near the carbon nanotube, their emission is quenched.
Pinals has tested this system with a number of proteins to identify which ones are more likely to bind to her sensor and kick off their DNA wrapping. Surprisingly, she found that while lots of protein was binding to the exposed surface of the SWNTs, relatively little DNA was being kicked off. She explains, “proteins first fill in the exposed carbon nanotube surface, potentially masking some of the DNA that’s there, but the sensor failure is mostly a function of biofouling.” In other words, it is nonselective, unpredictable proteins sticking to the surface and junking up the sensor that causes instability and inhibits fluorescence response. One way to overcome that barrier is by filling in any exposed nanotube surface with an anti-biofouling polymer such as polyethylene glycol (PEG). However, deterring protein adsorption while still allowing access for the analyte of interest to bind to the SWNT is a difficult balancing act.

While biofouling remains a frustrating issue, these results also open new opportunities in sensor development. Knowing what proteins preferentially bind to her SWNT sensors, Pinals has tried pre-passivating the SWNT surface with desired proteins that can detect other biomolecules of interest. By doing so, she is able to develop sensors that make use of the body’s natural biomolecular interactions rather than having to chemically engineer them from scratch. In addition to detecting molecules that are indicative of diseases like Huntington’s and Parkinson’s disease, tuning the protein corona can also help sensors target certain locations within the body while shielding them from destruction by the immune system.
Adapting Nanosensors to Detect SARS-COV-2
Since late spring, Pinals’ work has taken on new urgency as several members of the lab pivoted their research to develop a SWNT-based sensor for SARS-CoV-2. Widespread testing remains challenging in the U.S., driven largely by a shortage of tests and low throughput. The gold standard of testing, a polymerase chain reaction (PCR) assay, is time consuming and requires specialized instruments, personnel, and certain chemical reagents that are in limited supply. “Having a test that can quickly triage potential infection patients into a ‘probably yes’ versus ‘probably no’ category would be inherently useful, even if it’s not as sensitive as PCR,” explains Landry. These sensors would also have to function in biological fluids such as saliva and nasal fluid. Luckily, this is where Pinals’ understanding of corona formation comes into play.
“We know proteins adsorb onto our SWNT,” explains Pinals, “so we use this to our advantage to adsorb a protein that in turn has a known binding interaction with the SARS-COV-2 virus.” In particular, the lab is exploring two proteins – the ACE2 receptor that exists on the surface of many cell types, allowing the virus to infect the cell, and the SARS-COV-2 antibody, an immune response protein that binds to the virus and prevents it from entering a cell. Through a combination of covalent chemistry techniques and tuning of the protein corona, the lab ultimately aims to turn their SWNTs into rapid, real time detectors of active infections.
