Berkeley Lab Scientists Create First 3-D Model of a Protein Critical to Embryo Development
The first detailed and complete picture of a protein complex that is tied to human birth defects as well as the progression of many forms of cancer has been obtained by an international team of researchers led by scientists with the U.S. Department of Energy (DOE)’s Lawrence Berkeley National Laboratory (Berkeley Lab). Knowing the architecture of this protein, PRC2, for Polycomb Repressive Complex 2, should be a boon to its future use in the development of new and improved therapeutic drugs.
“We present a complete molecular organization of human PRC2 that offers an invaluable structural context for understanding all of the previous biochemical and functional data that has been collected on this complex,” says Berkeley Lab biophysicist Eva Nogales, an electron microscopy expert who led this research. “Our model should also be an invaluable tool for the design of new experiments aimed at asking detailed questions about the mechanisms that enable PRC2 to function and how those mechanisms might be exploited.”

Nogales, who holds joint appointments with Berkeley Lab, the University of California (UC) at Berkeley, and the Howard Hughes Medical Institute (HHMI), is one of two corresponding authors of a paper describing this research in the journal eLife. The paper is titled “Molecular Architecture of Human Polycomb Repressive Complex 2.” The other corresponding author is Claudio Ciferri, previously a postdoctoral fellow in the Nogales Lab, now with Novartis Vaccines and Diagnostic. Additional authors are Gabriel Lander, Alessio Maiolica, Franz Herzog and Ruedi Aebersold.
“PRC2 controls stem cell differentiation by regulating the expression of specific genes through the binding and methylation of histones, the proteins in chromatin that help bundle DNA into nucleosomes,” Nogales explains. “This is why PRC2 has been one of the top targets in drug development efforts by pharmaceutical companies.”
Despite the numerous biochemical and molecular studies that have been done on PRC2, little has been known about the protein’s overall architecture and the manner in which its different components interact to coordinate histone-binding and methylation. Nogales, Ciferri and their colleagues have closed this knowledge gap with their model, which they painstakingly constructed from data gathered via multiple sources including three-dimensional electron microscopy, mass spectrometry, protein biochemistry, crystal structure docking, and chemical cross-linking.
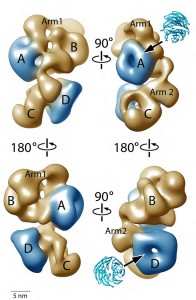
The 3D model they constructed features a structural reconstruction at 21 Angstroms and shows PRC2 with all its subunits and functional domains forming a complex with the protein called “AEBP2,” which serves as a stabilizing co-factor for PRC2. The PRC2-AEBP2 complex was shown to consist of four large lobes interconnected by two narrow arms. Clearly identified within this structure are regions where PRC2 and AEBP2 interact.
“This is a robust model that shows the overall architecture and domain organization of the PRC2-AEBP2 complex with unprecedented detail,” says Nogales. “From this model we can see how AEBP2 helps stabilize PRC2 and also how it plays a major role in coordinating the activity of different PRC2 subunits. Our model also provides us with a framework for understanding how various elements of the PRC2 complex involved in regulated histone modification can act either independently or synergistically in response to different chromatin environments.”

Not only does this new model put previous biochemical data on PRC2 in perspective, it also points the way to new testable hypotheses on how PRC2 interacts with chromatin that should inspire future research of PRC2 function and regulation. For example, from this model, Nogales, Ciferri and their co-authors have localized the molecular structures of individual domains involved in histone binding and methylation. The data suggests a mechanism as to how PRC2 might work in the chromatin environment.
“PRC2 is recruited at specific sites along the DNA that need to be silenced, aided by regulators such as AEBP2,” Nogales says. “Our domain localizations provide important information on how PRC2 can bind nucleosomes during the gene-silencing process. Such information can be used to guide specific deletion or cross-linking mutants for future drug design efforts.”
This research was supported by the National Institute of General Medical Sciences, the Howard Hughes Medical Institute and by funding from the European Union Seventh Framework Program PROSPECTS.
